Biostatistics Cluster Quickstart Guide
This guide will help you get your account, login, and run your first job.
- Requesting an account
- Logging in
- Copy scripts and data to the cluster
- Creating a batch script
- Submitting your job
- Checking the status of your job
Requesting an account
All members of the Biostatistics department are eligible to use the Biostatistics cluster for free. Users receive credits to run 20,000 CPU hours worth of computing each semester. As of Fall 2022, the Biostat cluster is no longer offering paid unlimited accounts. Users requiring computing beyond the free credits are encouraged to use Great Lakes. As of Fall 2019, all incoming Biostatistics students are automatically allocated a free cluster account. All other members of the Biostatistics department may request a user account by completing this form.
Accessing the Cluster
The Biostatistics cluster may be accessed from the command line or from a graphical web portal. New cluster users who are not familiar with using Linux from the command line or the SSH protocol are encouraged to start with the graphical web portal as it has a smaller learning curve. The cluster may only be accessed from oncampus or from the University VPN.
Graphical Web Portal
The cluster graphical web portal is accessible from any web browser. Simply navigate to biostat-login.sph.umich.edu and login with your University Credentials. After you log in, you will see the portal home page.
From the portal home page, you can select an application from the top menu bar. More on each application can be found below in this guide.
Command Line Linux
Connecting from a Windows computer
The following is an example of connecting to the cluster from a computer running the Windows operating system. This example uses PuTTY which can be downloaded here. Once PuTTY is installed and launched a window like this will appear. In the Host Name field you should enter: biostat-login.sph.umich.edu and then click Open.
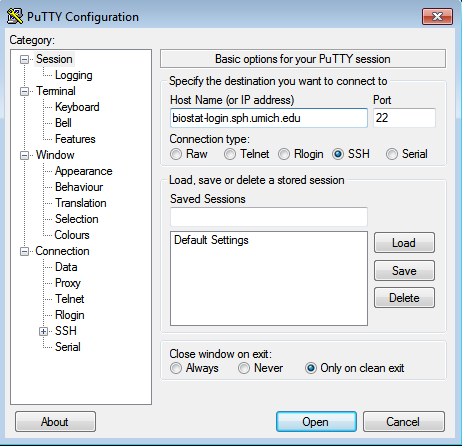
The first time you connect to the cluster you may see a warning. This is normal, click Yes.
A terminal will open where you will login using your uniqname, kerberos password and Duo two-factor. Note that as you type your password, nothing will appear on the password line. This is normal; just finish typing your password and hit "enter/return" on your keyboard.
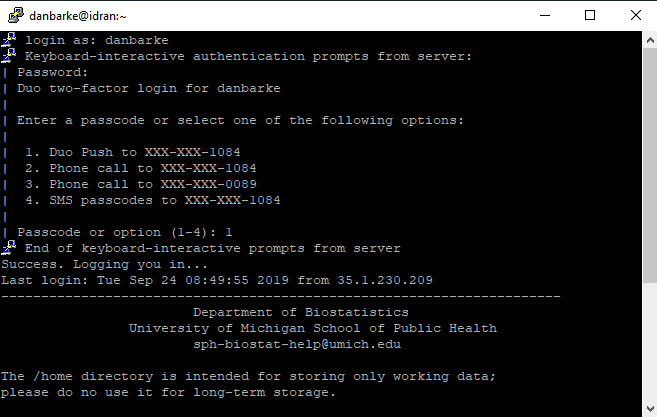
Connecting from a Mac or Linux Computer
The following is an example of connecting to the cluster from a computer running the MacOS or Linux operating systems. Both MacOS and Linux have an application called terminal. Once terminal is opened, the username that you want to connect as and the server that you wish to connect to must be specified.
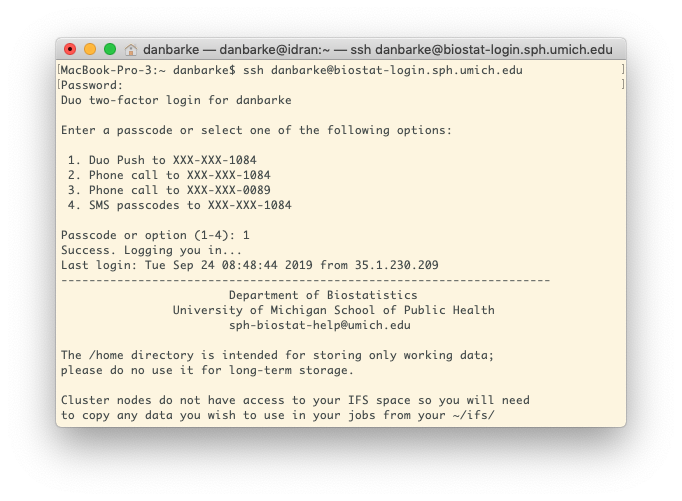
In the terminal window type ssh [email protected] and then hit enter/return on your keyboard. Replace uniqname with your University uniqname. You will be prompted for your University kerberos password. Note that as you type your password, nothing will appear on the password line. This is normal; just finish typing your password and hit "enter/return" on your keyboard. You will then be prompted to complete Duo two-factor authentication.
The following is an example of opening a session from a Mac or Linux computer
Copy scripts and data to the cluster
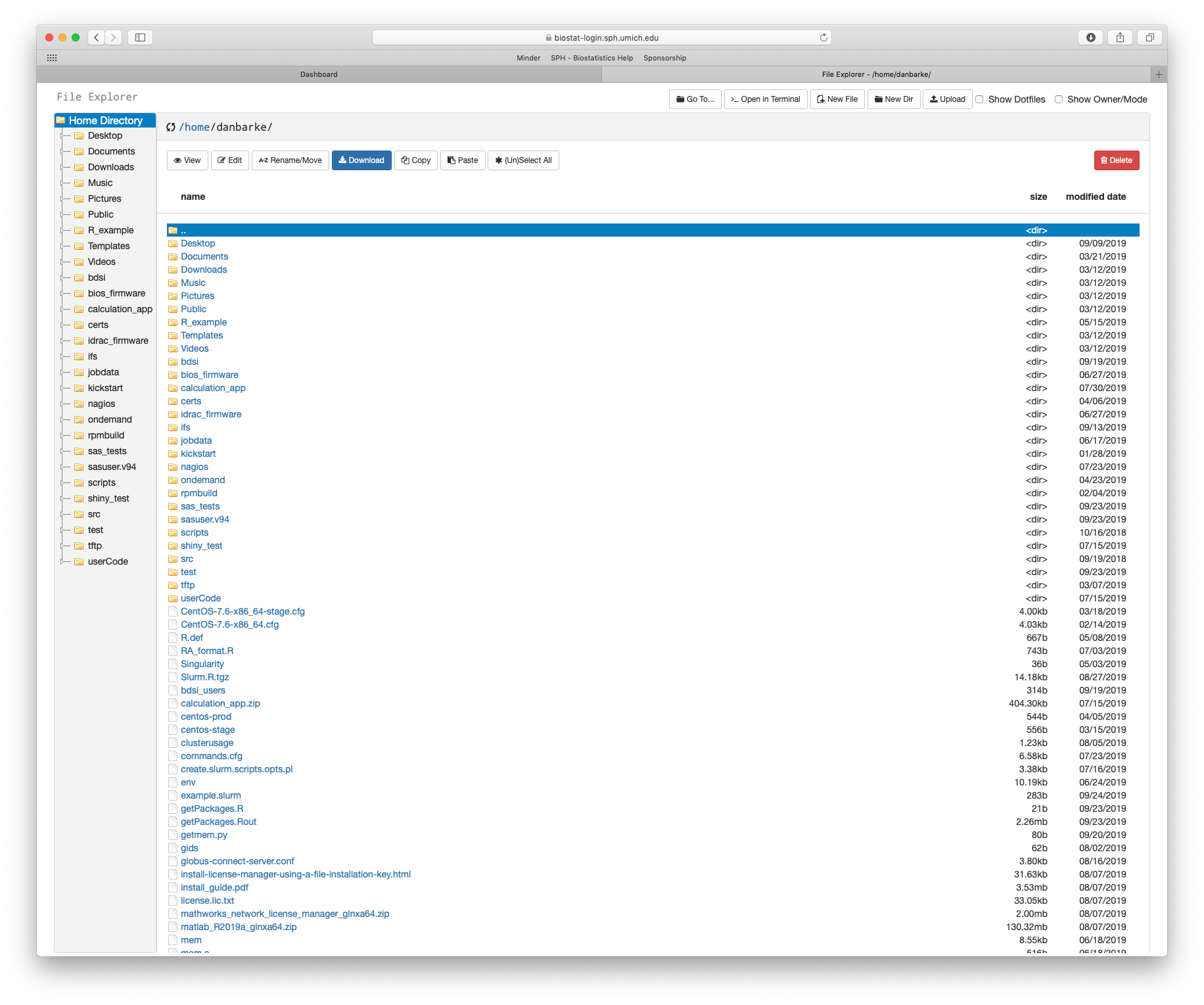
Your scripts and data files must be uploaded to the cluster in order to do computing with them. The easiest way to upload files to the cluster is through the "files" application of the graphical web portal. Simply navigate to biostat-login.sph.umich.edu and login with your University Credentials. After you log in, you will see the portal home page. On the top menu bar select "Files" and then "Home Directory". This will open a new tab in which you can use the File Explorer application. This application allows you to upload files to the cluster, download files from the cluster, view and edit files and directories. The buttons in the File Explorer application are named well and likely do not need more explanation.
Creating a batch script
A batch script is a specially formatted text file that specifies to the cluster the resources (CPUs, Memory, time) that the job will need. At the bottom of the batch script is a line that calls the script that should be executed. Batch scripts may be created locally on your computer and uploaded to the cluster, or they may be created directly on the cluster.
Here is a quick sample batch script for running a single job with a default resource allocation of 1 cpu core and 1000MB of memory running for 1 hour. It will execute an R script named script.R
#!/bin/bash
#SBATCH --job-name=hello_world
#SBATCH --time=1:00:00
#SBATCH [email protected]
#SBATCH --mail-type=END,FAIL,BEGIN
#SBATCH --mem=1000m
#SBATCH --cpus-per-task=1
R CMD BATCH --no-save --no-restore script.R
Submitting your job
Assuming you saved the above batch script to job.txt in your home directory you can submit the job with sbatch like this:
$ sbatch job.txt
You will see the job id for the job you just submitted and will receive an email when the job starts, when it ends, and if it should fail.
Checking the status of your job
You can check the status of your job in the queue with the squeue command:
$ sq
or
or
$ squeue -u $USER
Biostat Cluster Cheat Sheet
Download the Biostat Cluster Cheat Sheet here
Using tmux
You may find that your session on the cluster disconnects frequently due to a poor
internet connection or disconnections from the campus VPN client. This can be disruptive
to your work requiring that you re-login and reopen your applications before continuing
your work. To minimize disruptions and frustration, you can use an application called
tmux. tmux allows you to "attach" to a previous session and continue right where you
left off.
To use tmux, first log into the cluster like normal. Then before you start any work,
run the command "tmux". Now you will see a green bar at the bottom of your window.
This indicates that you are working within a tmux session.
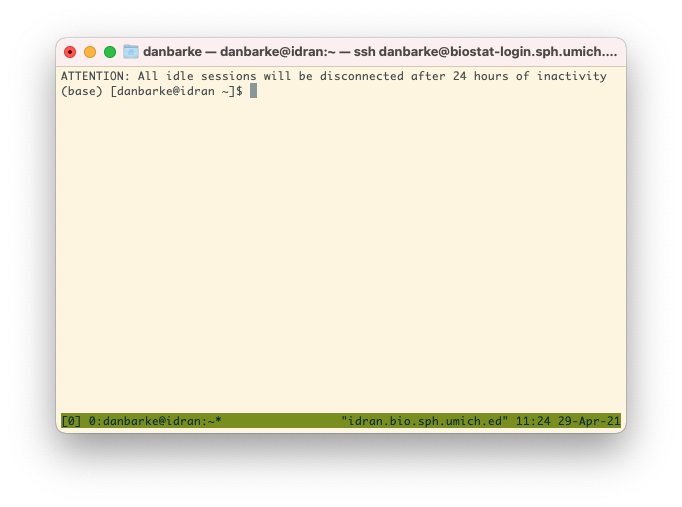
Go about your work as you normally would. If your session gets disconnected, you can
reconnect to this tmux session.
After a disconnect, log into the cluster again and then run "tmux attach". This will
reconnect you to your previous session. Continue working like normal. tmux has many
features to increase productivity, like multi-window and split screen support. You
can find more information about tmux here.


